User Tools
Sidebar
Add this page to your book
Remove this page from your book
This is an old revision of the document!
Table of Contents
Whole slide images and virtual slides
PMA.core supports a variety of image capturing techniques and modalities.
A full list of currently supported formats is available from our website at https://www.pathomation.com/formats.
You can read more about our philosophy regarding the support of vendor-specific file format in our press release.
Brightfield
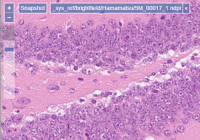
PMA.core slide reading engine supports over 35 different vendor-specific file formats that are used to encapsulate brightfield microscopy data. We offer a mix of open source community formats (OMERO TIFF & ZARR, [DICOM, …), as well as legacy ones like Olympus Webview.
A full up-to-date list is available on our website at https://www.pathomation.com/formats.
Fluorescence

DAPI, Cy3, Cy5, … If these terms sound familiar, changed are you're engaged in fluorescent microscopy. We've been working with many vendors throughout the years to optimally support this imaging type. You can toggle individual channels directly in PMA.core, and do histogram clipping as you see fit. Advanced downstream application (that run on top of PMA.core) like PMA.studio can even visualize the histogram of each channel for you, and apply additional per-channel gamma correction.
An up-to-date list of supported fluorescent file formats is available on our website at https://www.pathomation.com/formats.
Z-stacking

Both brightfield and fluorescent slides are often generated on a microtome, which results in smooth flat surface areas. However, when working with cytological samples, or doing microscopy in crop research, this is now always the case. Smears and similar materials are typically captured through z-stacking, whereby a sample is traversed in miniscule steps of one or two micron at the time.
An up-to-date list of z-stacking formats supported by PMA.core is available on our website at https://www.pathomation.com/formats.
Time series

